
・View per-atom labels for element, serial number, NMR shielding (when available).・Display formats: wire frame, tubes, ball & stick/bond type, space fill (CPK) style.・Manipulate multiple structures as an ensemble.・Use multiple synchronized or independent views of same structure (customizable).・View numeric value for any structural parameter.・Rotate, translate and zoom in 3D in any display using mouse operations and/or a precision positioning toolbar.At the end of each line, you must specify if the atom belongs to fragment 1 or fragment 2 with « 1 » or « 2 ».Features new to GaussView 6 are in blue features enhanced in GaussView 6 are in green.
Gaussian qst2 full#
Then:īefore the XYZ part, the numbers are for: charge of the full system, multiplicity of the full system, charge of fragment 1, multiplicity of fragment 1, charge of fragment 2, multiplicity of fragment 2. If have a iodine in your system, go here and choose the basis set for this element. Iodine is not defined in the 6-31G basis sets in Gaussian. Make a calculation with iodine and 6-31G type basis set

To use different basis sets for the atomsĭon’t forgot the 0 at the end of the list of atoms: # SCRF=(Solvent=Water) Guess=Read Geom=CheckPoint # IRC=(CalcFC,NoRaman,Reverse,MaxPoints=25)įor example, a first optimization in gas phase and then in water: After a few steps of IRC, finish by optimizing the geometry in a regular way. The « Forward » direction is going on the other side of the vibration. To do an IRC in the « Reverse » direction, see below. In this case, 60 steps of 1 degree for the csc3 coordinate will be made (so 61 values will be calculated), starting at 70 degres. Look for « modredundant » in the manual.Īnother option is presented below. #P M062X/6-31G* Opt=(CalcFC,NoRaman,TS,NoEigenTest)īelow, we do 25 steps of -0.1 Angstrom between atoms 19 and 24. Either you use « CalcFC », or you use a checkpoint file with « ReadFC » instead. The frequencies must have been computed before. If there are a lot of atoms to freeze, see below (look for « ReadOptimize » in the manual): This will fix the distance between atoms 3 and 4, the angle between atoms 5, 6 and 7 and the dihedral angle between atoms 6, 7, 8 and 9.
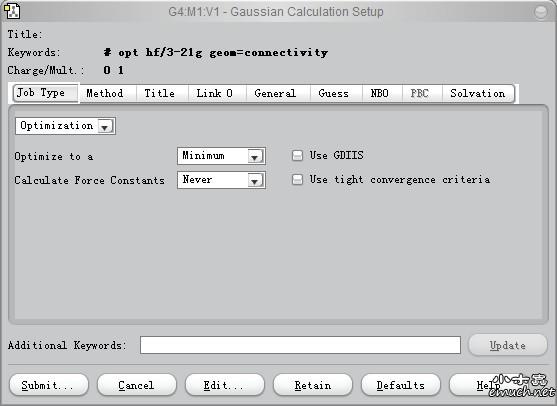
To optimize a geometry by fixing coordinates If you still have issues, check for the convergence criteria by simply looking for « YES » or « NO » in the output file.
Gaussian qst2 how to#
See below how to do two calculations in a row. Another option if you still have a problem is to optimize at a low level of theory and then use the level you want. If the SCF fails to converge, using SCF=XQC can be helpful. You can also compute the frequencies before the optimization with « Opt=(CalcFC,NoRaman) », or at every step with « Opt=(CalcAll) ». To decrease the size of the steps, « Opt=(MaxStep=5) » (default value is 30, and I sometimes even use 1). If this fails because of the number of cycles, you can use « Opt=(MaxCycle=300) ». For a simple optimization of the geometry If the calculation fails because of the size of the files, you can use at the beginning of the input file something like: %rwf=f1,2000Mb,f2,2000Mb,f3,2000Mb,f4,2000Mb,f5,-1. « Test » says this is a test calculation, and the results should not be send to the Gaussian archive. « #P » requests more details in the output file. « #T » at the beginning of the route requests minimal output from the program. I am only giving details which should help someone to know what to look at in the Gaussian manual. Otherwise, I give below a few tips to prepare input files for Gaussian09. There are also some tutorials online such as this one. My tutorial on « How to find a mechanism with Gaussian (and Opt’n Path) » explains how to optimize a structure, and then we focus on the transition state. Some webpages list frequent errors, such as this page. If you don’t find the information you need, you can look in the archives of CCL. If you are looking on how to use Gaussian, the best place to start is here.
